You are here
Jon's Blog
| Post date | ||
|---|---|---|
| Morpholino for antibody validation - example |
Webb AB, Lengyel IM, Jörg DJ, Valentin G, Jülicher F, Morelli LG, Oates AC. Persistence, period and precision of autonomous cellular oscillators from the zebrafish segmentation clock. Elife. 2016 Feb 13;5. pii: e08438. doi: 10.7554/eLife.08438. |
Friday, April 1, 2016 - 13:41 |
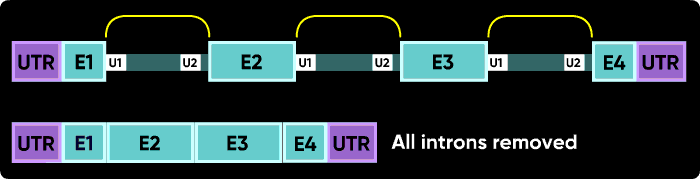
| Splicing outcomes: targeting for exon skipping or intron inclusion |
Someone asked me why some Morpholinos usually cause exon skips and others cause intron inclusions... U1 and U2 (or U11 and U12) snRNPs mark the positions on pre-mRNA of the splice junctions for the spliceosome. There is a U1 snRNP that binds in the intron near the e2i2 junction and a U2 snRNP that binds on the other side of the intron at the i2e3 junction. Figure 1 |
Wednesday, March 23, 2016 - 15:12 |
| RNA therapeutics: beyond RNA interference and antisense oligonucleotides |
Nice review, a few years old, describing Morpholinos and other antisense as therapeutics. RNA therapeutics: beyond RNA interference and antisense oligonucleotides. |
Tuesday, February 23, 2016 - 12:14 |
| A nucleic acid binder, poor at binding MO |
Sen D, Patel G, Patel SS. Homologous DNA strand exchange activity of the human mitochondrial DNA helicase TWINKLE. Nucl Acids Res. 2016;[Epub ahead of print] doi:10.1093/nar/gkw098 https://nar.oxfordjournals.org/content/early/2016/02/16/nar.gkw098.full DNA binds in the protein TWINKLE, but Morpholino is very poor at that binding. No surprise, it's likely due to lack of charge, but this is a nice published example of the low-binding characteristic. |
Wednesday, February 17, 2016 - 09:42 |
| Antisense oligonucleotide-directed inhibition of nonsense-mediated mRNA decay |
Here's an interesting technique: inhibition of nonsense-mediated decay with oligos. I don't think the antisense oligos were Morpholinos. Antisense oligonucleotide-directed inhibition of nonsense-mediated mRNA decay. http://www.nature.com/nbt/journal/vaop/ncurrent/full/nbt.3427.html |
Tuesday, February 2, 2016 - 10:24 |
| Use of Morpholinos to Regulate Gene Expression in the Brain |
Here is a review of tools to manipulate gene expression in the brain. Just before the Conclusion is the section "Use of Morpholinos to Regulate Gene Expression in the Brain". Walters BJ, Azam AB, Gillon CJ, Josselyn SA, Zovkic IB. Advanced In vivo Use of CRISPR/Cas9 and Anti-sense DNA Inhibition for Gene Manipulation in the Brain. Front Genet. 2016. doi:10.3389/fgene.2015.00362 http://journal.frontiersin.org/article/10.3389/fgene.2015.00362/full |
Monday, January 25, 2016 - 11:18 |
| Morpholino-driven gene editing: A new horizon for disease treatment and prevention |
Though I did in the early days of commercial Morpholinos, now I generally don't enter review articles into the Morpholino database. This one came along and I want to keep a record of it, particularly for the list of diseases in Table 1. So, it goes here. Subbotina E, Koganti SR, Hodgson-Zingman D, Zingman LV. Morpholino-driven gene editing: A new horizon for disease treatment and prevention. Clin Pharmacol Ther. 2015 Oct 16. doi: 10.1002/cpt.276. [Epub ahead of print] |
Thursday, December 17, 2015 - 13:44 |
| Example of splice enhancer targeting |
Liquori A, Vaché C, Baux D, Blanchet C, Hamel C, Malcolm S, Koenig M, Claustres M, Roux AF. Whole USH2A Gene Sequencing Identifies Several New Deep Intronic Mutations. Hum Mutat. 2015 Nov 2. doi: 10.1002/humu.22926. [Epub ahead of print] http://onlinelibrary.wiley.com/doi/10.1002/humu.22926/abstract |
Thursday, December 3, 2015 - 09:40 |
| Bathing zebrafish embryos with Vivo-Morpholinos |
Wong TT, Zohar Y. Production of reproductively sterile fish by a non-transgenic gene silencing technology. Sci Rep. 2015 Oct 29;5:15822. doi: 10.1038/srep15822. This group bathed zebrafish embryos (with chorion intact) in Vivo-Morpholino solutions, with oligos targeting dead-end (dnd). |
Monday, November 2, 2015 - 10:09 |
| Genetic compensation induced by deleterious mutations but not gene knockdowns |
When assessing gene function, gene knockdown reveals what can be hidden by compensation in mutants Rossi A, Kontarakis Z, Gerri C, Nolte H, Hölper S, Krüger M, Stainier DYR. Genetic compensation induced by deleterious mutations but not gene knockdowns. Nature. 2015;[Epub ahead of print] doi:10.1038/nature14580 |
Monday, July 13, 2015 - 11:15 |